17-10245
ChIPAb+ Histone H3.3 - ChIP Validated Antibody and Primer Set
from rabbit, purified by affinity chromatography
Synonym(s):
Histone H3.3
Sign Into View Organizational & Contract Pricing
All Photos(4)
About This Item
UNSPSC Code:
12352203
eCl@ss:
32160702
NACRES:
NA.32
Recommended Products
biological source
rabbit
Quality Level
clone
polyclonal
purified by
affinity chromatography
species reactivity
mouse, human
manufacturer/tradename
ChIPAb+
Upstate®
technique(s)
ChIP: suitable
dot blot: suitable
western blot: suitable
NCBI accession no.
UniProt accession no.
shipped in
dry ice
Gene Information
human ... H3F3B(3021)
General description
All ChIPAb+ antibodies are individually validated for chromatin precipitation, every lot, every time. Each ChIPAb+ antibody set includes control primers (tested every lot by qPCR) to biologically validate your IP results in a locus-specific context. The qPCR protocol and primer sequences are provided, allowing researchers to validate ChIP protocols when using our antibody in their chromatin context. Each set also includes a negative control antibody to ensure specificity of the ChIP reaction.
The ChIPAb+ Histone H3.3 set includes the Histone H3.3 antibody, a Normal rabbit IgG, and control primers which amplify a 87 bp region of ChIP Primers, GAPDH Coding Region D2 Human. The Histone H3.3 and negative controls are supplied in a scalable "per ChIP" reaction size and can be used to functionally validate the precipitation of Histone H3.3 -associated chromatin.
The ChIPAb+ Histone H3.3 set includes the Histone H3.3 antibody, a Normal rabbit IgG, and control primers which amplify a 87 bp region of ChIP Primers, GAPDH Coding Region D2 Human. The Histone H3.3 and negative controls are supplied in a scalable "per ChIP" reaction size and can be used to functionally validate the precipitation of Histone H3.3 -associated chromatin.
Specificity
Broad species cross-reactivity expected based on 100% sequence homology.
This antibody recognizes human Histone H3.
Immunogen
Epitope: a.a. 85-105
KLH-conjugated linear peptide corresponding to human Histone H3.3.
Application
Chromatin Immunoprecipitation:
Representative lot data.
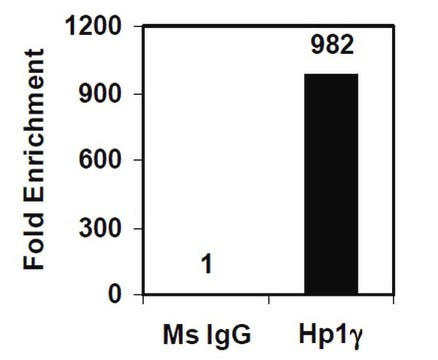
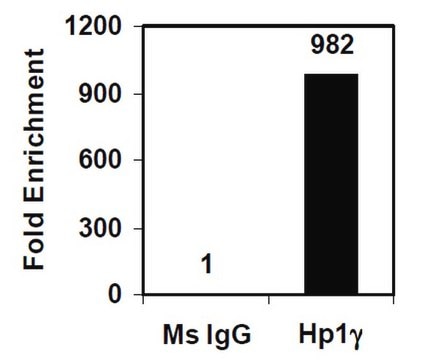
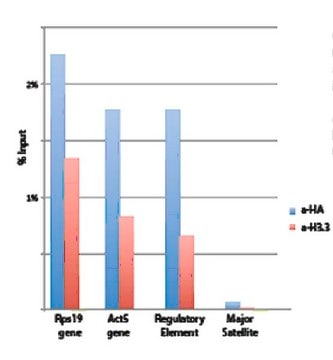
Sonicated chromatin prepared from HeLa cells (1 X 10E6 cell equivalents per IP) were subjected to chromatin immunoprecipitation using 2 µg of either Normal rabbit IgG or 2 µg Anti-Histone H3.3 and the Magna ChIP A Kit (Cat. # 17-610). Successful immunoprecipitation of Histone H3.3 associated DNA fragments was verified by qPCR using ChIP Primers, GAPDH Coding Region D2 as a positive locus, and beta-actin promoter primers as a negative locus. (Figure 2). Data is presented as percent input of each IP sample relative to input chromatin for each amplicon and ChIP sample as indicated.
Please refer to the EZ-Magna ChIP A (Cat. # 17-408) or EZ-ChIP (Cat. # 17-371) protocol for experimental details.
Western Blot Analysis:
Representative lot data.
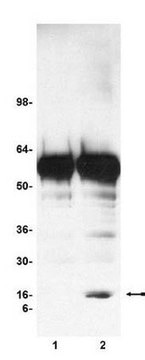
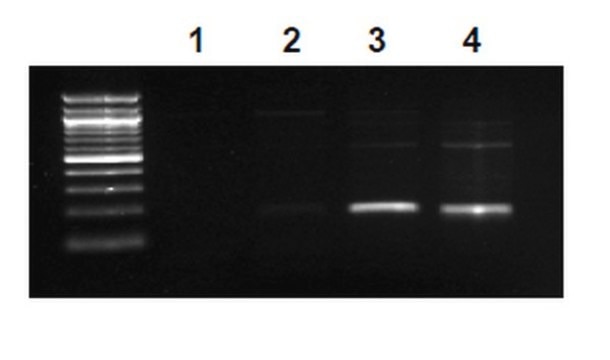
HeLa acid extract was probed with Anti-Histone H3.3 (1 µg/mL). Proteins were visualized using a Donkey Anti-Rabbit IgG secondary antibody conjugated to HRP and a chemiluminescence detection system.
Arrow indicates Histone H3.3 (~17 kDa). (Figure 3).
Dot Blot Analysis;
Representative lot data.
Recombinant H3.2 and H3.3 spotted on PVDF (1, 10, 100, 1,000 ng histone per slot). Detection with 0.5 µg/mL a-H3.3 antibody in TBS-T, 5% milk. (Figure 4).
Image courtesy of Simon Elsässer, Laboratory of David Allis, Rockefeller University, New York.
Representative lot data.
Sonicated chromatin prepared from HeLa cells (1 X 10E6 cell equivalents per IP) were subjected to chromatin immunoprecipitation using 2 µg of either Normal rabbit IgG or 2 µg Anti-Histone H3.3 and the Magna ChIP A Kit (Cat. # 17-610). Successful immunoprecipitation of Histone H3.3 associated DNA fragments was verified by qPCR using ChIP Primers, GAPDH Coding Region D2 as a positive locus, and beta-actin promoter primers as a negative locus. (Figure 2). Data is presented as percent input of each IP sample relative to input chromatin for each amplicon and ChIP sample as indicated.
Please refer to the EZ-Magna ChIP A (Cat. # 17-408) or EZ-ChIP (Cat. # 17-371) protocol for experimental details.
Western Blot Analysis:
Representative lot data.
HeLa acid extract was probed with Anti-Histone H3.3 (1 µg/mL). Proteins were visualized using a Donkey Anti-Rabbit IgG secondary antibody conjugated to HRP and a chemiluminescence detection system.
Arrow indicates Histone H3.3 (~17 kDa). (Figure 3).
Dot Blot Analysis;
Representative lot data.
Recombinant H3.2 and H3.3 spotted on PVDF (1, 10, 100, 1,000 ng histone per slot). Detection with 0.5 µg/mL a-H3.3 antibody in TBS-T, 5% milk. (Figure 4).
Image courtesy of Simon Elsässer, Laboratory of David Allis, Rockefeller University, New York.
Research Category
Epigenetics & Nuclear Function
Epigenetics & Nuclear Function
Research Sub Category
Histones
Histones
This ChIPAb+ Histone H3.3 -ChIP Validated Antibody & Primer Set conveniently includes the antibody & the specific control PCR primers.
Packaging
25 assays per set. Recommended use: ~2 μg of antibody per chromatin immunoprecipitation (dependent upon biological context).
Quality
Chromatin Immunoprecipitation:
Representative lot data.
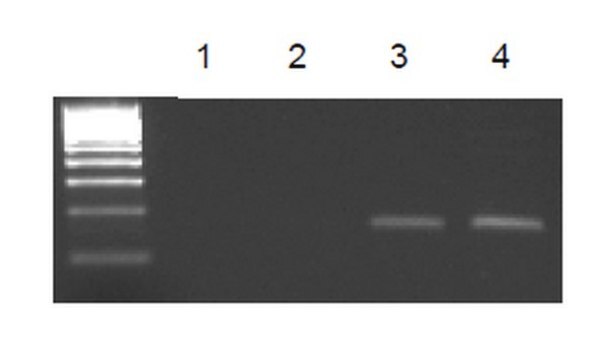
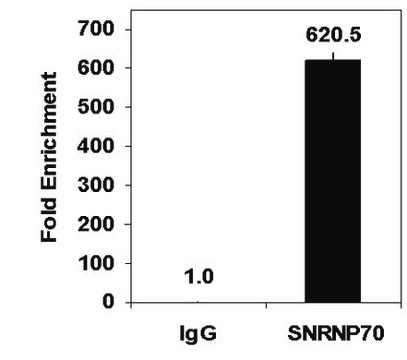
Sonicated chromatin prepared from HeLa cells (1 X 10E6 cell equivalents per IP) were subjected to chromatin immunoprecipitation using 2 µg of either Normal Rabbit IgG or 2 µg of Anti-Histone H3.3 and the Magna ChIP® A Kit (Cat. # 17-610). Successful immunoprecipitation of Histone H3.3 associated DNA fragments was verified by qPCR using ChIP Primers, GAPDH Coding Region D2 (Figure 1).
Please refer to the EZ-Magna ChIP A (Cat. # 17-408) or EZ-ChIP (Cat. # 17-371) protocol for experimental details.
Representative lot data.
Sonicated chromatin prepared from HeLa cells (1 X 10E6 cell equivalents per IP) were subjected to chromatin immunoprecipitation using 2 µg of either Normal Rabbit IgG or 2 µg of Anti-Histone H3.3 and the Magna ChIP® A Kit (Cat. # 17-610). Successful immunoprecipitation of Histone H3.3 associated DNA fragments was verified by qPCR using ChIP Primers, GAPDH Coding Region D2 (Figure 1).
Please refer to the EZ-Magna ChIP A (Cat. # 17-408) or EZ-ChIP (Cat. # 17-371) protocol for experimental details.
Target description
~17 kDa observed
Physical form
Affinity purified
Anti-Histone H3.3 (rabbit polyclonal). One vial containing 50 µg of purified rabbit polyclonal in buffer containing 0.1M Tris-Glycine (pH 7.4), 150 mM NaCl with 0.05% sodium azide before the addition of glycerol to 30%. Store at -20° C.
Concentration: 0.7 mg/mL
Normal Rabbit IgG. One vial containing 125 µg of Rabbit IgG in 125 µL of storage buffer containing 0.05% sodium azide. Store at -20°C.
Control Primers, GAPDH Coding Region D2 Human. One vial containing 75 μL of 5 μM of each primer specific for human GAPDH coding region. Store at -20°C.
FOR: GCC ATG TAG ACC CCT TGA AGA G
REV: ACT GGT TGA GCA CAG GGT ACT TTA T
Concentration: 0.7 mg/mL
Normal Rabbit IgG. One vial containing 125 µg of Rabbit IgG in 125 µL of storage buffer containing 0.05% sodium azide. Store at -20°C.
Control Primers, GAPDH Coding Region D2 Human. One vial containing 75 μL of 5 μM of each primer specific for human GAPDH coding region. Store at -20°C.
FOR: GCC ATG TAG ACC CCT TGA AGA G
REV: ACT GGT TGA GCA CAG GGT ACT TTA T
Storage and Stability
Stable for 1 year at -20°C from date of receipt. Handling Recommendations: Upon first thaw, and prior to removing the cap, centrifuge the vial and gently mix the solution. Aliquot into microcentrifuge tubes and store at -20°C. Avoid repeated freeze/thaw cycles, which may damage IgG and affect product performance.
Note: Variability in freezer temperatures below -20°C may cause glycerol containing solutions to become frozen during storage.
Note: Variability in freezer temperatures below -20°C may cause glycerol containing solutions to become frozen during storage.
Analysis Note
Control
Includes normal rabbit IgG and primers specific for human GAPDH coding region.
Includes normal rabbit IgG and primers specific for human GAPDH coding region.
Other Notes
Concentration: Please refer to the Certificate of Analysis for the lot-specific concentration.
Legal Information
MAGNA CHIP is a registered trademark of Merck KGaA, Darmstadt, Germany
UPSTATE is a registered trademark of Merck KGaA, Darmstadt, Germany
Disclaimer
Unless otherwise stated in our catalog or other company documentation accompanying the product(s), our products are intended for research use only and are not to be used for any other purpose, which includes but is not limited to, unauthorized commercial uses, in vitro diagnostic uses, ex vivo or in vivo therapeutic uses or any type of consumption or application to humans or animals.
wgk_germany
WGK 2
flash_point_f
Not applicable
flash_point_c
Not applicable
Certificates of Analysis (COA)
Search for Certificates of Analysis (COA) by entering the products Lot/Batch Number. Lot and Batch Numbers can be found on a product’s label following the words ‘Lot’ or ‘Batch’.
Already Own This Product?
Find documentation for the products that you have recently purchased in the Document Library.
Analysis of neonatal brain lacking ATRX or MeCP2 reveals changes in nucleosome density, CTCF binding and chromatin looping.
Kernohan, KD; Vernimmen, D; Gloor, GB; Berube, NG
Nucleic Acids Research null
William Chang et al.
Epigenetics & chromatin, 16(1), 4-4 (2023-01-27)
Cellular differentiation is marked by temporally and spatially coordinated gene expression regulated at multiple levels. DNA methylation represents a universal mechanism to control chromatin organization and its accessibility. Cytosine methylation of CpG dinucleotides regulates binding of methylation-sensitive DNA-binding transcription factors
Joseph Aaron Goldman et al.
Developmental cell, 40(4), 392-404 (2017-03-02)
Chromatin regulation is a principal mechanism governing animal development, yet it is unclear to what extent structural changes in chromatin underlie tissue regeneration. Non-mammalian vertebrates such as zebrafish activate cardiomyocyte (CM) division after tissue damage to regenerate lost heart muscle.
Liquan Zhou et al.
Development (Cambridge, England), 144(3), 519-528 (2016-12-21)
During development from oocyte to embryo, genetic programs in mouse germ cells are reshaped by chromatin remodeling to orchestrate the onset of development. Epigenetic modifications of specific amino acid residues of core histones and their isoforms can dramatically alter activation
Renee J Tamming et al.
Cell reports, 31(13), 107838-107838 (2020-07-02)
ATRX gene mutations have been identified in syndromic and non-syndromic intellectual disabilities in humans. ATRX is known to maintain genomic stability in neuroprogenitor cells, but its function in differentiated neurons and memory processes remains largely unresolved. Here, we show that
Our team of scientists has experience in all areas of research including Life Science, Material Science, Chemical Synthesis, Chromatography, Analytical and many others.
Contact Technical Service