SRP0125
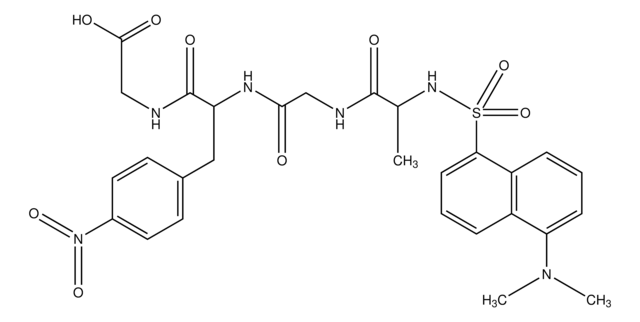
LSD1 substrate (Di-methylated K4_H3)
≥90% (HPLC), aqueous solution
Synonym(s):
HIST1H3E, Histone H3.1
About This Item
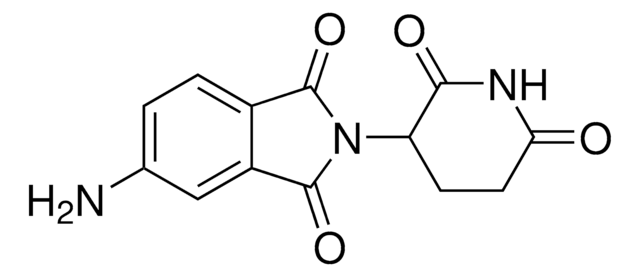
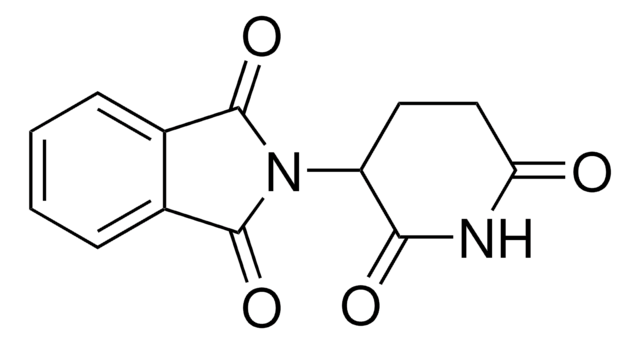
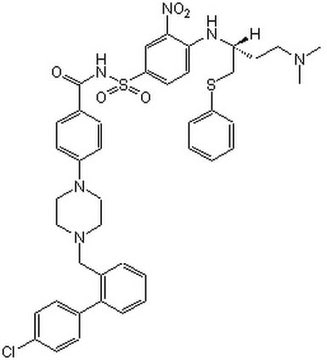
Recommended Products
product name
LSD1 substrate (Di-methylated K4_H3), ≥90% (HPLC)
biological source
human
assay
≥90% (HPLC)
form
aqueous solution
mol wt
2283 Da
packaging
pkg of 500UL
concentration
0.55 mg/mL
UniProt accession no.
shipped in
dry ice
storage temp.
−70°C
Gene Information
human ... HIST1H3E(8353)
General description
Application
Biochem/physiol Actions
Physical form
Preparation Note
flash_point_f
Not applicable
flash_point_c
Not applicable
Certificates of Analysis (COA)
Search for Certificates of Analysis (COA) by entering the products Lot/Batch Number. Lot and Batch Numbers can be found on a product’s label following the words ‘Lot’ or ‘Batch’.
Already Own This Product?
Find documentation for the products that you have recently purchased in the Document Library.
Articles
Epigenetic modifications are thought to occur through two key interconnected processes—DNA methylation and the covalent modification of histones.
Our team of scientists has experience in all areas of research including Life Science, Material Science, Chemical Synthesis, Chromatography, Analytical and many others.
Contact Technical Service