MAC4L
MetaPolyzyme
lyophilized powder
Synonym(s):
Multilytic Enzyme Mix
About This Item
Recommended Products
form
lyophilized powder
Quality Level
technique(s)
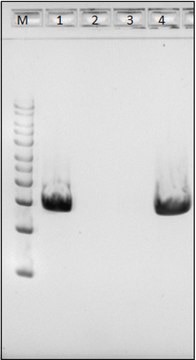
DNA extraction: suitable
suitability
suitable for microbiology
application(s)
microbiology
shipped in
wet ice
storage temp.
−20°C
Related Categories
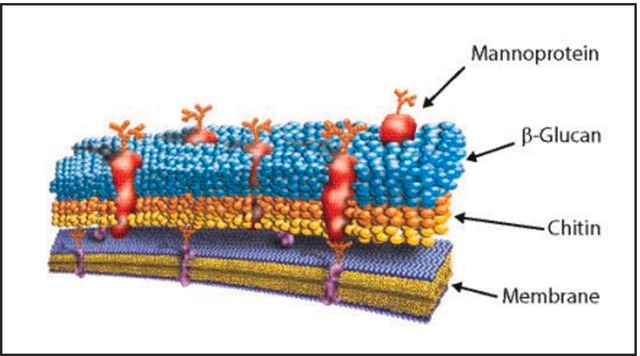
General description
Application
Biochem/physiol Actions
Components
- Achromopeptidase
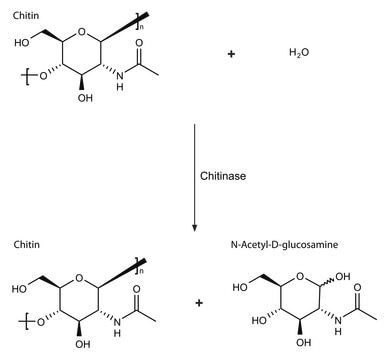
- Chitinase
- Lyticase
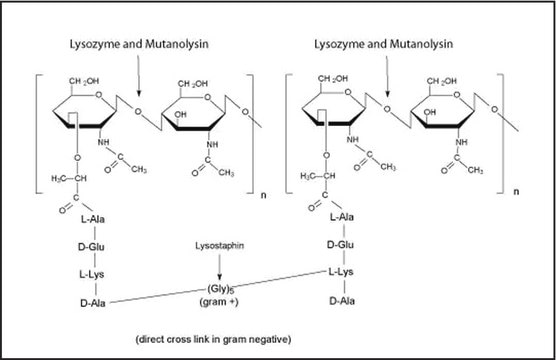
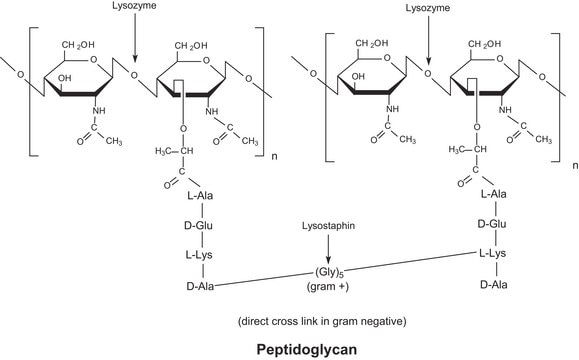
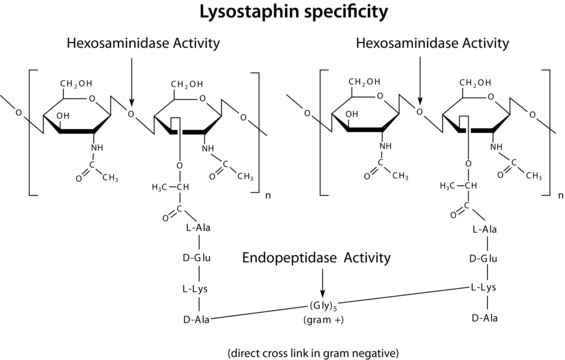
- Lysostaphin
- Lysozyme
- Mutanolysin
signalword
Danger
hcodes
pcodes
Hazard Classifications
Resp. Sens. 1
Storage Class
11 - Combustible Solids
wgk_germany
WGK 3
flash_point_f
Not applicable
flash_point_c
Not applicable
Certificates of Analysis (COA)
Search for Certificates of Analysis (COA) by entering the products Lot/Batch Number. Lot and Batch Numbers can be found on a product’s label following the words ‘Lot’ or ‘Batch’.
Already Own This Product?
Find documentation for the products that you have recently purchased in the Document Library.
Customers Also Viewed
Articles
Enzymes provide a non-mechanical method for cell lysis and protoplast preparation. It may seem like a simple process to throw in your enzyme, stick your tube in the water-bath and walk away, but what is actually going on in that process?
An overview of human microbiome research, workflow challenges, sequencing, library production, data analysis, and available microbiome reagents to support your research.
Our team of scientists has experience in all areas of research including Life Science, Material Science, Chemical Synthesis, Chromatography, Analytical and many others.
Contact Technical Service